Innovation translates
Our lab combines biomedical imaging, systems biology, and data visualization to explore six key areas: cancer research, chronic kidney disease, regenerative medicine, fibrosis, aging and brain health, and neurodegeneration.
Theme 1
Decoding drug resistance in cancer treatments
Our research uncovered signaling networks and discovered cell-cycle-driven protein interactomics behind Tyrosine Kinase Inhibitor (TKI) inhibitors in non-small lung cancer
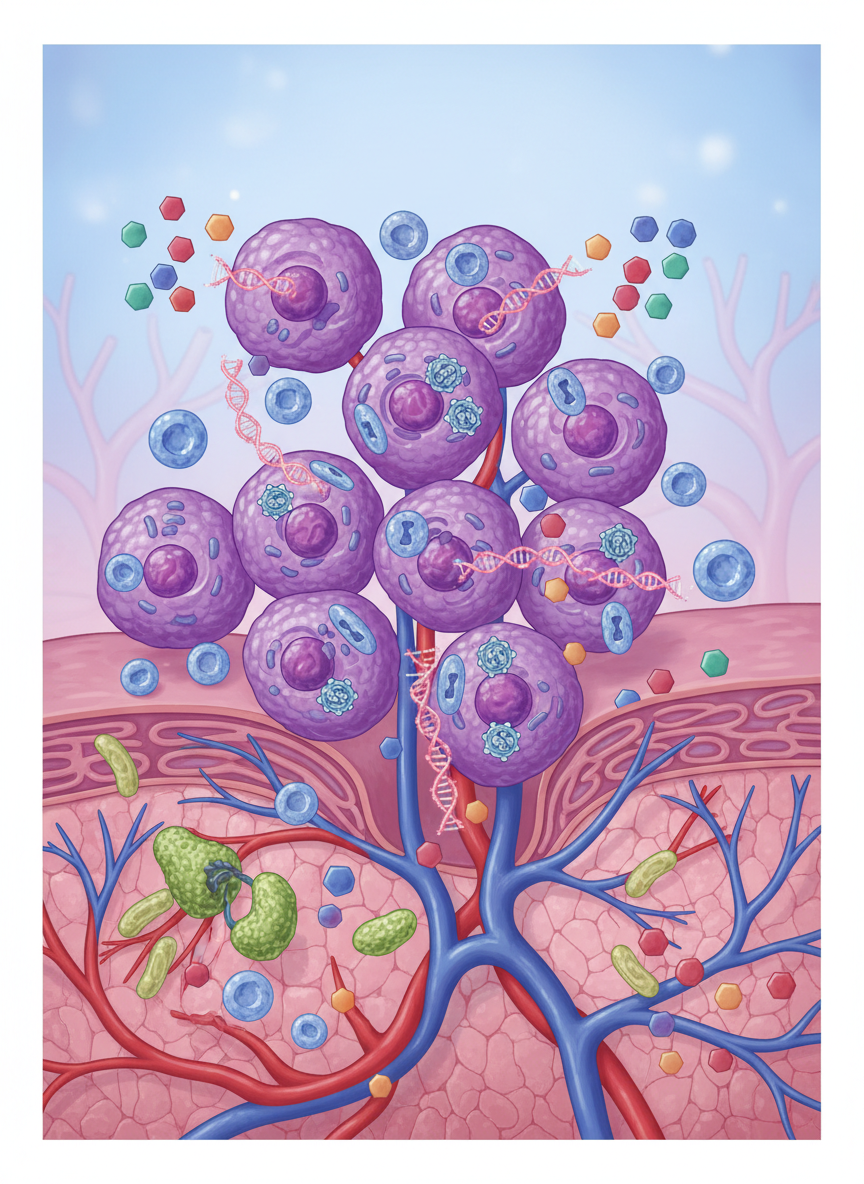
Theme 2
Predicting stem cell function in regenerative medicine
Our lab developed a digital spatial transcriptomics-based metric to model stromal-immune cell communication for predicting efficacy in stem cell therapies
Theme 3
Uncovering immune senescence in aging
Our lab envisions single cell dissection of senescence signatures in a multi-omics fashion to quantify biological senescence beyond chronogical aging

Enabling Spatial Biomarkers
We advance cancer, immunity, aging, and chronic diseases through single-cell analysis, spatial omics, and integrative data visualization to uncover mechanisms and accelerate therapies.